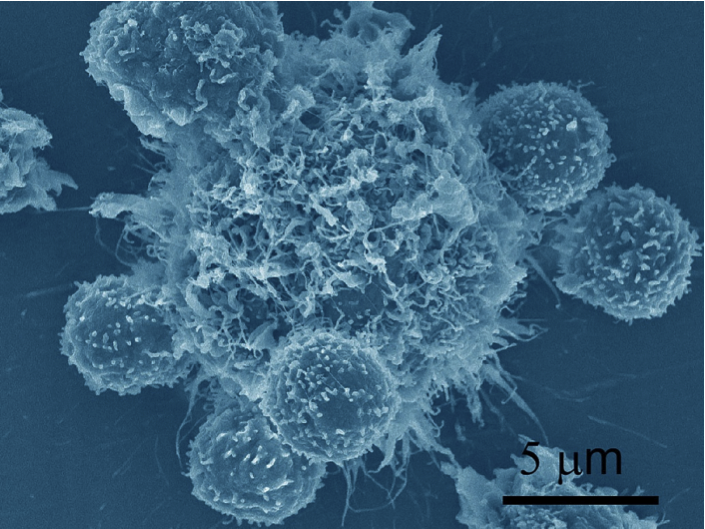
Circulating tumor cells (CTCs) are shed into the bloodstream from both primary and metastatic tumors. Some CTCs will play a vital role as metastatic precursors for the blood-borne spread of cancer. Novel microfluidic-based CTC capture technology developed at MGH has enabled the detection and isolation of this very rare cell with high captured cell purity and yield. Performance testing of the CTC-iChip using blood samples from patients with many different types of cancer demonstrates that this platform isolates a high number of CTCs per blood sample, has the ability to capture CTCs from essentially all patients tested, and the capacity to dynamically monitor CTC quantity in a clinically meaningful way in real time. This new non-invasive testing (or “liquid biopsy” based on CTC analysis) has the potential to transform the approaches and clinical decision-making for the detection, monitoring, and treatment of cancer.
Read MoreOver the past decade, bioengineers from the MGH Center for Engineering in Medicine have overcome numerous technical hurdles in identifying, sorting, and capturing rare cells like CTCs in a process that preserves their viability. The capability of the CTC-iChip to capture the rare CTCs sparked the interests of MGH Cancer Center clinicians who recognize the potential for CTC-based diagnostics and therapeutics could be realized only if these rare cells were captured in high numbers with high purity, and under conditions that make the cells compatible for downstream analysis using both standard clinical diagnostic cytopathology as well as advanced molecular and genomic testing. The MGH team’s test results indicate that the high capture rates and viable condition of the cells isolated via the CTC-iChip make it an ideal tool for detecting CTCs from patients with early stage cancer, determining responsiveness to cancer treatment, and identifying early cancer recurrence or metastasis. Read more about the engineering design process of the CTC-iChip.
Studying the CTCs captured using the CTC-iChip technology
Enumeration of single CTCs and identification of CTC microclusters
The microfluidic CTC capture platform, now known as the CTC-iChip, can identify, sort, and capture CTCs in whole blood samples (Nagrath et al, 2007). CTCs have been successfully isolated from patients with many different cancers including lung, prostate, pancreas, breast, colon, melanoma, and brain. Evaluation of the clinical utility of CTC enumeration or counting showed that the presence of detectable CTCs in the blood served as an independent prognostic factor in patients with cancers of the breast, prostate, and colon (Yu et al, 2011). Analysis of baseline (pretreatment) CTCs numbers among different cancer patients were not necessarily correlated with standard measures of tumor mass, including tumor size detected by CAT scan or serum protein markers like PSA, prostate-specific antigen (Stott et al, STM, 2010). These findings of absolute CTC counts and pretreatment CTC numbers derived from performance testing of the CTC-Chip are consistent with those of other cancer researchers using different techniques for measuring CTCs. Most scientists believe that the number of CTCs in the blood is not simply a measure of tumor volume, but instead, it may reflect additional biological features, possibly including tumor vascularity or invasiveness, both of which may have distinct prognostic contributions.

Immunofluorescence image of a glioblastoma (GBM) CTC alongside a WBC isolated from blood of a patient with GBM.
The first reported identification of brain tumor CTCs within the peripheral blood of patients with glioblastoma (GBM) was made possible using the MGH team’s CTC-iChip in negative depletion mode (Sullivan et al, 2014). Although GBM is the most common and aggressive primary malignant brain tumor, it is locally invasive within the brain and rarely metastasizes to distant organs. The investigators tested for the presence of GBM CTCs and showed that GBM sheds CTCs with invasive mesenchymal characteristics into the blood circulation. In a pilot study of CTCs from a rare patient with metastatic disease, the metastatic GBM CTCs were primarily mesenchymal and showed additional (acquired) mutations absent in the primary tumor. The identification of GBM CTCs and their detailed characterization may provide new insight into the invasive properties of these aggressive brain tumors, ultimately identifying new therapeutic opportunities to suppress proliferation of primary GBMs.
Detection of CTC microclusters. A surprising insight derived from the performance testing of the CTC-iChip has been the discovery of CTC microclusters within the circulation of patients with known cancer. CTC clusters of 3 to 15 cells in several prostate and lung cancer patient samples were first identified using the “second generation” herringbone CTC-Chip, which appeared to preserve multicellular clusters (Stott et al, PNAS, 2010). Interestingly, CTC clusters had never been identified using the “first generation” micropost CTC-Chip in which tight distances between the microposts presumably prevented the passage of large cellular clusters. In subsequent studies testing the negative depletion mode of the CTC-iChip, the capture platform identified clusters of 2 to 6 CTCs in breast and pancreatic cancers, consistent with the team’s previous detection of CTC clusters. CTC clusters were further examined in patients with metastatic breast cancer and mouse breast cancer models (Aceto et al, 2014). In patients with breast cancer, single-cell resolution RNA sequencing of CTC clusters and single CTCs, matched with individual blood samples, identified the cell junction protein plakoglobin as highly differentially expressed. In patients, both abundance of CTC clusters and high tumor plakoglobin denoted adverse outcomes. These CTC clusters held together through plakoglobin-dependent intercellular adhesion, although rare, greatly contribute to the metastatic spread of cancer. The CTC-Chip studies isolating CTCs clusters in breast (Ozkumur et al, 2013, Yu et al, 2013, Aceto et al, 2014), pancreatic (Ozkumur et al, 2013), lung (Stott et al, PNAS, 2010), and prostate (Stott et al, PNAS, 2010) cancer patients and mouse models of cancer point to CTC clusters as critical mediators of cancer metastasis.
Serial measurements of CTCs to monitor response to therapy
CTC-iChip testing demonstrates that the capture platform is sensitive enough to correlate CTC numbers over time with response to therapy. The studies have shown that the number of CTCs in patients with lung, prostate, and other cancers declines rapidly after the initiation of effective chemotherapy, hormonal therapy, or targeted kinase inhibition (Nagrath et al, 2007, Maheswaran et al, 2008, Stott et al, STM, 2010). Drug treatment-induced changes in CTC numbers within individual patients followed over time were well correlated with standard measures of tumor response, irrespective of baseline CTC counts.
Monitoring the decline in CTC numbers after curative surgical resection of localized prostate cancer identified patients who were at risk for disease occurrence. In a pilot study of 19 patients with localized prostate cancer, CTCs were detected in eight cases. In most of these cases, CTCs dropped to undetectable levels within 24 hours of surgical resection in contrast to a few cases wherein a more delayed decline in CTC counts after surgery was evident. None of these patients developed a recurrence at a one-year follow up. Investigators postulated that localized cancers may invade the vasculature and shed CTCs before metastases are established, a concept that provides hope for early cancer detection applications. In this small study, the investigators were able to estimate the half-life of CTCs in the blood circulation after surgical resection of the primary prostate tumor. The very rapid postoperative decline in CTC counts in a subset of patients indicates that an individual CTC is likely to be present in the blood for less than 24 hours, and if all sources of tumor shedding have been eliminated, the number of CTCs in blood should rapidly drop (Stott et al, STM, 2010).
In captured CTCs from a genetically engineered mouse model of melanoma, the investigators analyzed CTC counts over time and demonstrated that CTC numbers rise as the primary tumor enlarges and metastases appear. Similarly, a brisk decline in CTCs concomitant with tumor shrinkage in response to targeted therapy with a selective drug was evident. In four patients undergoing drug therapy for melanoma, CTC numbers were generally correlated with clinical evaluation of disease status (Luo et al, 2014).
Molecular and genomic characterization of individual CTCs
Beyond CTC enumeration, the high purity and high yield of the captured cells permits not only global CTC expression analysis, but also identification of expression patterns and signaling pathways within individual CTCs. This important information may help clinicians to better understand the unique molecular and genomic abnormalities that may drive a patient’s cancer diagnosis and response to first or second line therapy. Genetic abnormalities not only define a cancer at diagnosis, but mutations over time can lead to acquired resistance to chemotherapy drugs and failure of drug therapy. Strategies based on evolving tumor mutational profiles and drug sensitivity patterns in individual patients are likely to become an essential component of “precision medicine” in oncology.
Identification of molecular aberrations in CTCs

Serial monitoring of EGFR genotypes revealed the emergence of additional activating EGFR mutations in CTCs during therapy
A cancer tumor can undergo molecular evolution as it shrinks, for example, in response to the administration of chemotherapy drugs. In a complex process involving clonal evolution and tumor heterogeneity, the tumor may acquire resistance to drug therapy. Noninvasive monitoring of the tumor genotype using CTC analysis during the course of treatment can provide important early information about the genetic evolution of tumors.
In serial measurements of lung CTCs, the investigators identified epidermal growth factor receptor (EGFR) mutations in CTCs from patients with non-small-cell lung cancer and these EGFR mutations in CTCs were consistent with the mutations known to be present in the primary tumor (from tissue biopsy) in 12 of 13 cases. During the long-term follow up of these patients, serial monitoring of CTCs from patients receiving EGFR tyrosine kinase inhibitor drug therapy showed the emergence of a secondary EGFR mutation associated with the development of clinical drug resistance. Furthermore, CTC analysis after prolonged drug therapy showed the acquisition of additional EGFR mutations that were below detection in the primary tumor biopsy (Maheswaran et al, 2008).
The use of CTCs to test for molecular (genetic) evolution of tumors during the course of treatment has also been applied in a study of prostate cancer, wherein the investigators demonstrated the presence of a tumor-specific chromosomal translocation – a fusion of two prostate cancer-specific genes that are normally separated – which occurs in about one-half of advanced prostate cancers. This preliminary study demonstrated that serial analysis of tumor-derived samples may allow real-time monitoring of the molecular evolution among tumor cells (Stott et al, STM, 2010).
In a proof-of-concept study, the investigators established long-term, patient-derived oligoclonal CTC cultures from CTCs isolated from six patients with metastatic estrogen receptor-positive breast cancer as one approach to study the molecular mechanisms behind breast cancer progression. Genomic sequencing of the individual CTC lines showed pre-existing mutations and newly acquired mutations. The CTC lines were tested for sensitivity to panels of single drug and drug combinations, including standard clinical regimens, as well as experimental agents targeting specific mutations, revealing potential new therapeutic targets. With optimization of CTC culture conditions, this strategy may help identify the best therapies for individual cancer patients over the course of their disease (Yu et al, 2014).
Evaluation of cell proliferation, cell signaling, and apoptosis
The cancer disease process requires cell division, growth, and survival, all of which is accomplished using extraordinarily complex systems of cell signaling networks or pathways that inform the cell whether to grow, proliferate, migrate, or die. In cancer research, mouse models, including transgenic mouse models, have been utilized for decades to improve the understanding of the disease process in patients. Mouse and human CTCs captured with the CTC-iChip are being analyzed to help our investigators characterize known key signaling networks as well as potential new signaling mechanisms that may be involved in the cancer cell’s uncontrolled growth and invasion of surrounding tissues.
Early pilot studies characterizing prostate CTCs from patients with local and metastatic prostate cancer revealed significant variation in the amount of cell proliferation among patients at different states of disease (Stott et al, STM, 2010). CTCs from metastatic prostate cancer patients who were responding to hormone withdrawal therapy showed a low proliferative index in contrast to treatment-resistant patients whose CTCs demonstrated a much higher proliferative score. Prior to this CTC study, limited information had been available about the types of tumor cells constituting CTCs and the extent to which these are representative of the proliferative or quiescent components of either primary or metastatic tumor deposits. In a study of androgen receptor (AR) readouts from CTCs captured before and after first-line androgen deprivation therapy, the findings revealed a profound switch from “AR-on” to “AR-off” CTCs populations in patients with castration-resistant prostate cancer. Measuring treatment-induced signaling responses within CTCs may help guide therapy in various types of cancer (Miyamoto et al, 2012).
Some CTCs are cells in the process of epithelial-mesenchymal transition (EMT), a state wherein adherent epithelial cells change to a migratory mesenchymal state, which has been implicated in metastasis. The technical challenges in analyzing EMT in CTCs have been overcome with the new tumor antigen-independent CTC-iChip, which not only can isolate CTCs from epithelial tumors, but also from nonepithelial cancers like melanoma and from cancer that has undergone EMT and lost all detectable epithelial expression like triple-negative breast cancer (Ozkumur et al, 2013). Using the CTC-iChip to investigate the role of EMT in human cancer, the investigators observed rare primary breast tumor cells simultaneously expressing mesenchymal and epithelial markers, but mesenchymal cells were highly enriched in the CTCs. Serial monitoring of CTCs in 11 breast cancer patients suggested an association of mesenchymal CTCs with disease progression. Mesenchymal cells occurred as both single cells and CTC clusters, expressing known EMT regulators, supporting a role for EMT in the blood-borne dissemination of human breast cancer (Yu et al, 2013).
Pancreatic cancer is an especially lethal cancer, which stems from the rapid dissemination of tumor cells leading to widespread metastasis. An initial characterization of mouse CTCs using partially purified CTCs identified activation of the Wnt signaling pathway as a recurrent event, potentially contributing to the circulating pancreatic epithelial cells being resistant to anoikis, an apoptopic (programmed cell death) pathway that helps cancer cell survive in the circulation (Yu et al, 2012). Extension of these studies to include single-cell RNA sequencing of human and mouse pancreatic CTCs captured using the CTC-iChip showed an unexpectedly high expression of stromal-derived extracellular matrix (ECM) proteins, including one protein that when removed from mouse cancer cells suppresses cell migration and invasiveness (Ting et al, 2014).
The future of the CTC test
The MGH team of bioengineers, molecular biologists, and cancer clinicians have designed, developed, tested, and applied novel microfluidic strategies to isolate and characterize CTCs in the bloodstream of patients with cancer. The innovation team has produced critical technological developments to efficiently isolate rare CTCs in suspension without the bias of tumor antigen selection, combined with the ability to perform high-fidelity single-cell RNA sequencing. Working in partnership with Johnson & Johnson, the MGH researchers hope to speed production of the CTC-iChip for expanded testing in larger cohorts of cancer patients to translate this research innovation to the cancer patient’s bedside.
Relevant publications
Nagrath S, Sequist LV, Maheswaran S, Bell DW, Irimia D, Ulkus L, et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature. 2007 Dec 20;450(7173):1235-9. PubMed PMID: 18097410; PubMed Central PMCID: PMC3090667
Maheswaran S, Sequist LV, Nagrath S, Ulkus L, Brannigan B, Collura CV, et al. Detection of mutations in EGFR in circulating lung-cancer cells. N Engl J Med. 2008 Jul 24;359(4):366-77. PubMed PMID: 18596266; PubMed Central PMCID: PMC3551471
Sequist LV, Nagrath S, Toner M, Haber DA, Lynch TJ. The CTC-chip: an exciting new tool to detect circulating tumor cells in lung cancer patients. J Thorac Oncol. 2009 Mar;4(3):281-3. PubMed PMID: 19247082
Stott SL, Lee RJ, Nagrath S, Yu M, Miyamoto DT, Ulkus L, et al. Isolation and characterization of circulating tumor cells from patients with localized and metastatic prostate cancer. Sci Transl Med. 2010 Mar 31;2(25):25ra23. PubMed PMID: 20424012; PubMed Central PMCID: PMC3141292
Stott SL, Hsu CH, Tsukrov DI, Yu M, Miyamoto DT, Waltman BA, et al. Isolation of circulating tumor cells using a microvortex-generating herringbone-chip. Proc Natl Acad Sci U S A. 2010 Oct 26;107(43):18392-7. PubMed PMID: 20930119; PubMed Central PMCID: PMC2972993
Yu M, Stott S, Toner M, Maheswaran S, Haber DA. Circulating tumor cells: approaches to isolation and characterization. J Cell Biol. 2011 Feb 7;192(3):373-82. Review. PubMed PMID: 21300848; PubMed Central PMCID: PMC3101098
Yu M, Ting DT, Stott SL, Wittner BS, Ozsolak F, Paul S, et al. RNA sequencing of pancreatic circulating tumour cells implicates WNT signaling in metastasis. Nature. 2012 Jul 26;487(7408):510-3. Erratum in: Nature. 2012 Oct 25;490(7421):570. PubMed PMID: 22763454; PubMed Central PMCID: PMC3408856
Miyamoto DT, Lee RJ, Stott SL, Ting DT, Wittner BS, Ulman M, et al. Androgen receptor signaling in circulating tumor cells as a marker of hormonally responsive prostate cancer. Cancer Discov. 2012 Nov;2(11):995-1003. PubMed PMID: 23093251; PubMed Central PMCID: PMC3508523
Yu M, Bardia A, Wittner BS, Stott SL, Smas ME, Ting DT, et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science. 2013 Feb 1;339(6119):580-4. PubMed PMID:23372014; PubMed Central PMCID: PMC3760262
Ozkumur E, Shah AM, Ciciliano JC, Emmink BL, Miyamoto DT, Brachtel E, et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Sci Transl Med. 2013 Apr 3;5(179):179ra47. PubMed PMID: 23552373; PubMed Central PMCID: PMC3760275
Luo X, Mitra D, Sullivan RJ, Wittner BS, Kimura AM, Pan S, et al. Isolation and molecular characterization of circulating melanoma cells. Cell Rep. 2014 May 8;7(3):645-53. PubMed PMID: 24746818; PubMed Central PMCID: PMC4079008
Yu M, Bardia A, Aceto N, Bersani F, Madden MW, Donaldson MC, et al. Cancer therapy. Ex vivo culture of circulating breast tumor cells for individualized testing of drug susceptibility. Science. 2014 Jul 11;345(6193):216-20. PubMed PMID: 25013076
Aceto N, Bardia A, Miyamoto DT, Donaldson MC, Wittner BS, Spencer JA, et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell. 2014 Aug 28;158(5):1110-22. PubMed PMID: 25171411; PubMed Central PMCID: PMC4149753
Ting DT, Wittner BS, Ligorio M, Vincent Jordan N, Shah AM, Miyamoto DT, et al. Single-cell RNA sequencing identifies extracellular matrix gene expression by pancreatic circulating tumor cells. Cell Rep. 2014 Sep 25;8(6):1905-18. PubMed PMID: 25242334; PubMed Central PMCID: PMC4230325
Sullivan JP, Nahed BV, Madden MW, Oliveira SM, Springer S, Bhere D, et al. Brain tumor cells in circulation are enriched for mesenchymal gene expression. Cancer Discov. 2014 Nov;4(11):1299-309. PubMed PMID: 25139148; PubMed Central PMCID: PMC4221467
Contact
| Daniel A. Haber, MD, PhD | Mehmet Toner, PhD | Shannon L. Stott, PhD | Shyamala Maheswaran, PhD | |
| 617-726-7805 | 617-724-5336 | 617-643-2658 | 617-724-6552 |




